The Rosetta energy function has its own idea about the "types" of atoms. (That is, which atoms are similar in how they behave in the Rosetta energy function.) While there are similarities in the atom typing scheme that Rosetta uses and those of other packages, there are differences.
Certain inputs in Rosetta also refer to atom types, often to allow for ambiguous representations. (For example, the Enzdes/Matcher constraint file format allows you to specify contraints based on atom types rather than atom names, to allow ambiguoius specification: In fullatom mode, the two carboxylate oxygens of ASP ('OD1' and 'OD2') are both typed 'OOC', and the sidechain nitrogen of ASN ('ND2') and GLN ('NE2') have the Rosetta type 'NH2O'.
Determining Rosetta AtomTypes
The atom types for Rosetta are specified in the "params file" for that residue. For standard residue types (e.g. the amino acids), those params files are located in the Rosetta database under main/database/chemical/residue_type_sets/.
The atom types are specified on the ATOM lines:
ATOM N Nbb NH1 -0.47 -0.350
ATOM CA CAbb CT1 0.07 0.100
ATOM C CObb C 0.51 0.550
ATOM O OCbb O -0.51 -0.550
The first column after the ATOM specifier is the atom name, and the second column (the Nbb/CAbb/etc.) is the Rosetta atom type. The third column (NH1/CT1/C/O) is the CHARMM molecular mechanics atom type, which is not used by default in Rosetta, but is read by certain specialty energy terms (the "MM" terms). (The fourth and fifth columns are partial charges.)
Full Atom
Under full atom mode, all atoms are present.
Amino Acids


(The original ChemDraw files for these figures are present in the images directory, if you wish to adjust them.)
Small molecules and other compounds
Rosetta atoms types are centered around and parameterized for representing amino acid residues. In representing non-protein residues, the protein-centric residue types are often repurposed based on heuristics of the small molecule structure.
Examine the params file for your small molecule for details on how the atoms are typed.
Centroid
Centroid mode uses a "United Atom" model, where apolar hydrogens are merged with their attached heavy atom. (Though polar hydrogens are preserved for calculating hydrogen bonds.)
Additionally, the protein sidechains are merged into a single 'CEN' "super atom".
Amino Acids
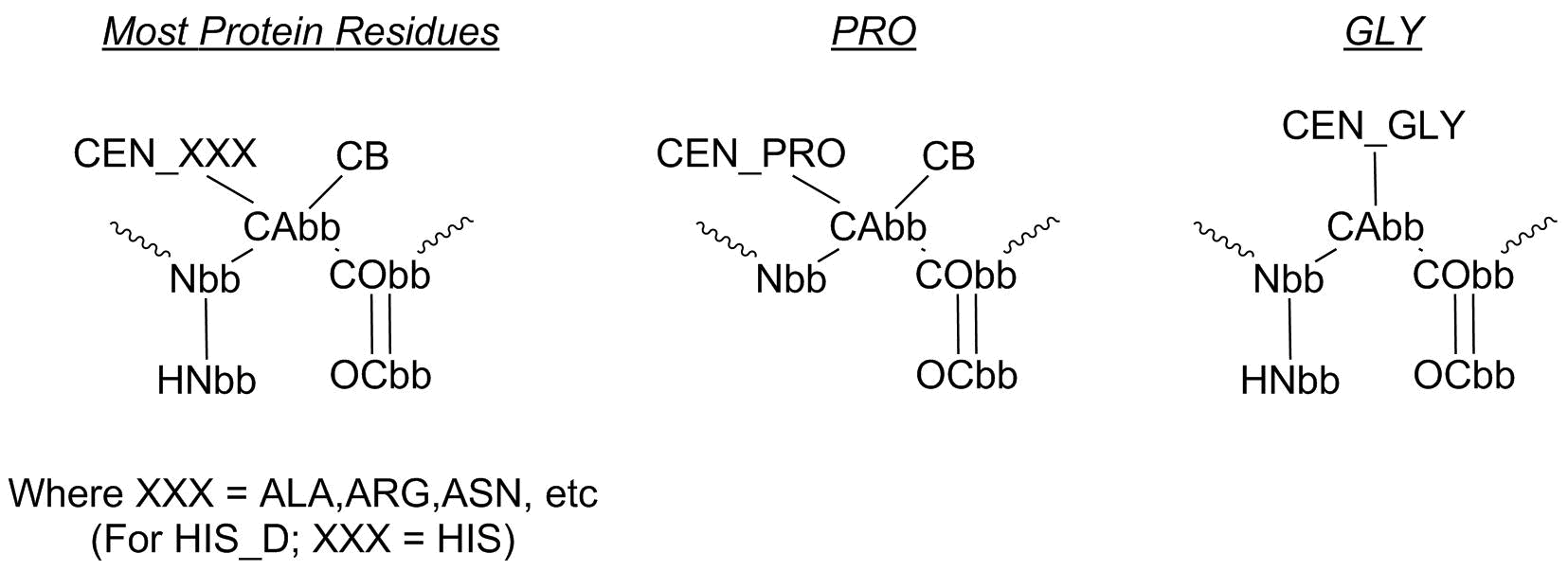
(The original ChemDraw files for this figure is present in the images directory, if you wish to adjust it.)
See Also
- Glossary: Brief definitions of Rosetta terms
- Rosetta overview: Overview of major concepts in Rosetta
- Score functions and score terms
- Solving a Biological Problem: Guide for applying Rosetta to your biological problem